Nam Tran, Ph.D., HCLD (ABB), FAACC
Stuart Cohen, M.D.
Sarah Waldman, M.D.
Larissa May, M.D.
Introduction
The current pandemic has created significant interest in COVID-19 related laboratory testing. Given the rapidly evolving testing landscape, it has become challenging to keep up with the numerous assays available on the market. Some assays have been marketed with questionable performance in recent weeks, while reagent shortages have forced many hospital laboratories to adopt multiple testing modalities to keep up with demand – creating additional educational challenges when describing COVID-19 testing.1,2 The goal of this blog article is to clarify the current state of testing for COVID-19 and provide recommendations for best practices. Information presented below is current at the time of posting this article.
SARS-CoV-2 Diagnostic Testing
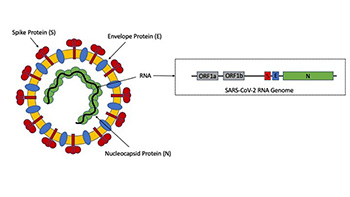
As of May 28, 2020, molecular and antigen testing are the only techniques capable of detecting the SARS-CoV-2 virus. Briefly, SARS-CoV-2 is an enveloped positive sense RNA virus. Molecular approaches are designed to target one or more genes (Figure 1).3 Common target genes include the envelope (E), nucleocapsid (N), spike protein (S), and the open reading frame (ORF) region. Some assays may target multiple regions within a gene, such as the assay used by the Center for Disease Control and Prevention (CDC). Molecular methods are primarily based on reverse transcriptase (RT) polymerase chain reaction (PCR). Other methods do exist but are less common. Early data (late 2019) suggested RT-PCR SARS-CoV-2 assays having a false negative rate of about 30%.4 Presently, the primary UC Davis assay has a sensitivity of greater than 97.5%, specificity of 100%, and can detect << 50 copies/mL of virus. On average, a typical nasopharyngeal swab sample from a symptomatic patient carries about 106 copies/mL of virus – several orders of magnitude above the detection limit modern PCR assays.5 False negatives are largely attributed to variable viral shedding of COVID-19 and/or bad specimen collection technique or processing. Specimen type also influences sensitivity, with nasopharyngeal swabs, nasal and mid-turbinate still being preferred for testing of symptomatic individuals.6 At UC Davis NP swabs are accepted with other specimen types currently under validation. Testing from lower respiratory tract specimens, such as bronchoalveolar fluid, are presently performed through Sacramento County Public Health Lab.
Point-of-care testing is also available, however current methods relying on isothermal methods exhibit poor sensitivity for SARS-CoV-2.7 Recent reports indicate these methods have up to 15% false negatives, and therefore, not considered appropriate for screening asymptomatic individuals. Any negative result would require confirmation by RT-PCR.
More recently, SARS-CoV-2 viral antigen testing has become available and could also be used at the point of care. Instead of detecting the viral RNA, antigen testing employs immunoassay techniques for detecting specific viral proteins.8 The benefits associated with antigen testing is speed (~15 minutes) and cost-effectiveness. Unfortunately, viral antigen kinetics following infection remains unknown. It is believed that SARS-CoV-2 antigens can be found shortly after RNA is detectable. Manufacturer data suggests a sensitivity of about 80% and specificity of 100% based on remnant nasopharyngeal swab viral transport media samples (n = 143) as well as prospectively collected samples (n = 48). More data are needed before widespread adoption of SARS-CoV-2 antigen testing can be made. To this end, molecular approaches remain the primary means for detecting SARS-CoV-2.
Serology Testing
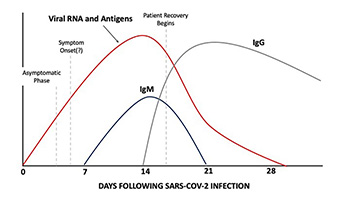
Serology testing measures the host antibody response following infection.9 SARS-CoV-2 serology testing is also based on immunoassay techniques and measures immunoglobulins (Ig) such as IgM, IgA, or IgG. Both IgA and IgM appear early and are less specific, while IgG is produced later and is more specific.10 IgG antibody levels may correlate with potential immunity (Figure 2). Data suggests IgM production begins as early as three days post-infection with IgG appearing as early as seven days post-infection. By day 14, the majority of individuals should have a detectable antibody response. Some, but not all, serology assays experience false positives due to previous exposure to non-SARS-CoV-2 human coronaviruses. Additionally, false negative serology results may occur if testing is performed too soon after exposure where antibody production is non-detectable (“window period”). To this end, it's important to note that positive SARS-CoV-2 serology results alone are not diagnostic, nor does it definitively indicate that a patient has had prior SARS-CoV-2 infection.1 It is also important to understand that not all antibodies produced against a virus, such as SARS-CoV-2, confer immunity. At present, no serology assay to date can differentiate between non-neutralizing and neutralizing antibodies, and it is not known how long we maintain immunity (if any) following SARS-CoV-2 infection.
The current UC Davis serology assay targets IgG antibodies against the S1/S2 domains of the spike protein and exhibits a sensitivity of 97.6% and specificity of 99.3%. Serology assay specificity is critical to success for COVID-19 due to the low disease prevalence impacting the positive predictive value of the test.1 Exclusion of IgM and IgA testing is due to the lack of specificity as well as more recent data showing IgG often rapidly rising in parallel to IgM following SARS-CoV-2 infection.10 Total immunoglobin tests (IgA, IgM, and IgG) have also emerged claiming the ability to detect high affinity antibodies to enhance sensitivity, while maintaining acceptable specificity.12 More studies are needed to compare the performance of all these assays under real world conditions. Regardless, it is recommended that COVID-19 serology1: (a) results should be interpreted in the context of the expected predictive values, positive and negative, (b) testing used to support the diagnosis of COVID-19 in patients presenting within 9 to 14 days of symptom onset in conjunction with molecular testing, (c) testing used to support observational epidemiological studies evaluating the prevalence of disease, or (d) testing used to help establish a diagnosis when patients present with late complications of COVID-19 illness, such as multisystem inflammatory syndrome in children. SARS-CoV-2 serology SHOULD NOT be used alone as a COVID-19 diagnostic test without paired molecular diagnostics, to determine immune status post-COVID-19, guide personal protective equipment use, or returning to the workplace decisions. Commercially available “rapid” serology tests (lateral flow assays) are not recommended for any setting due to significant false negative and false positive rates.
Other Chemistry Tests
In addition to the testing approaches above, routine biochemical testing can aid the diagnosis and management of COVID-19.13 Procalcitonin (PCT) is presently used as a biomarker of bacterial infection and for antimicrobial stewardship. During a viral infection, interferon gamma production inhibits PCT release, thus COVID-19 patients should present with normal PCT levels. Although true for the majority of patients, some recent studies suggest PCT values may be elevated in individuals with severe COVID-19. This observation may be due to the presence of bacterial co-infection, and/or tissue injury-mediated release of PCT due to COVID-19 related tissue damage. Interleukin-6 (IL-6) is another biochemical marker under discussion for COVID-19. Non-surviving COVID-19 patients presented with significantly higher IL-6 compared to survivors. Unfortunately, IL-6 is presently available as a referral laboratory test and should only be used if the result would prompt any management change. Lastly, coagulation testing may help identify patients at risk for COVID-19 related coagulopathy. D-dimer values are significantly higher in patients with severe COVID-19.
References
- Centers for Disease Control and Prevention COVID-19 Serology Guidelines: Accessed on May 28, 2020
- Infectious Disease Society of America Primer: May 4, 2020.
- Wu F, et al. A new coronavirus associated with human respiratory disease in China. Nature 2020;579:265-269.
- Wang W, et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 2020;323:1843-1844.
- Wolfel R, et al. Virologic assessment of hospitalized patients with COVID-19. Nature 2020;581;465-469.
- Infectious Disease Society of America Guidelines for the Treatment and Management of Patients with COVID-19 (Updated May 6, 2020): Accessed on May 28, 2020.
- United States Food and Drug Administration Press Release (May 14, 2020): Accessed on May 28, 2020
- Quidel Sophia 2 SARS Antigen FIA product insert: Accessed on May 28, 2020.
- American Society for Microbiology Communications (May 19, 2020): Accessed on May 28, 2020.
- Long QX, et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nature Medicine 2020.
- Diasorin LIAISON SARS-CoV-2 S1/S2 IgG product insert (EN – 200/007-798, 03 – 2020-5).
- Roche Diagnostics SARS-CoV-2 serology assay: Accessed on May 28, 2020.
- International Federation for Clinical Chemistry (IFCC) Guide on COVID-19 (May 25, 2020): Accessed on May 28, 2020.



