Software developed at UC Davis analyzes calcium ‘sparks’ that can contribute to arrhythmia
SparkMaster 2 will allow biomedical researchers to analyze normal and abnormal calcium activity in human cells
A team of UC Davis and University of Oxford researchers have developed an innovative tool: SparkMaster 2. The open-source software allows scientists to analyze normal and abnormal calcium signals in cells automatically.

Calcium is a key signaling molecule in all cells, including muscles like the heart. The new software enables the automatic analysis of distinct patterns of calcium release in cells. This includes calcium "sparks," microscopic releases of calcium within cardiac cells associated with irregular heartbeats, also known as arrhythmia.
A research article demonstrating the capabilities of SparkMaster 2 was published in Circulation Research.
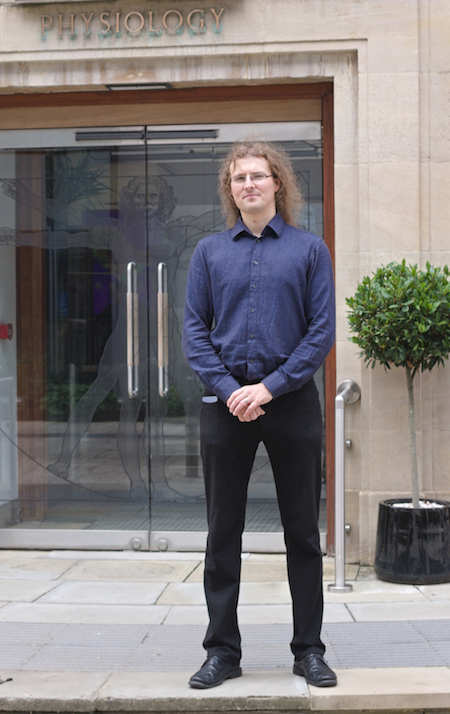
Jakub Tomek, the first author of the research article, is a Sir Henry Wellcome Fellow in the Department of Physiology, Anatomy and Genetics at the University of Oxford. He spent his fellowship year at UC Davis, working with Distinguished Professor Donald M. Bers.
"It was great to present SparkMaster 2 at recent conferences and see the enthusiastic response. I felt it would be an outlier and that few people would care. But many people were excited about having a new analysis tool that overcomes many of the limitations they have experienced with prior tools," Tomek said.
Fellowship at UC Davis leads to updated tool
Problems with how and when calcium is released by cells can have an impact on a range of diseases, including arrhythmia and hypertension. To understand the mechanisms behind these diseases, researchers use fluorescent calcium indicators and microscopic imaging that can measure the calcium changes at the cellular level.
However, the resulting data files are large and challenging to analyze by hand, so customized software analysis tools are essential.
At the Bers Lab at UC Davis, Tomek discussed analysis tools with Christopher Y. Ko, an assistant project scientist. They identified the need for better, more user-friendly software, which led to the development of SparkMaster 2.
The new tool builds upon the success of SparkMaster, which was released in 2007 by Bers and Eckard Picht. Picht is a physician-scientist who worked with Bers, first at Loyola University Chicago and then for a few years at UC Davis. At the time, there was no user-friendly free software that could analyze calcium sparks — a crucial part of their research — so they developed one.
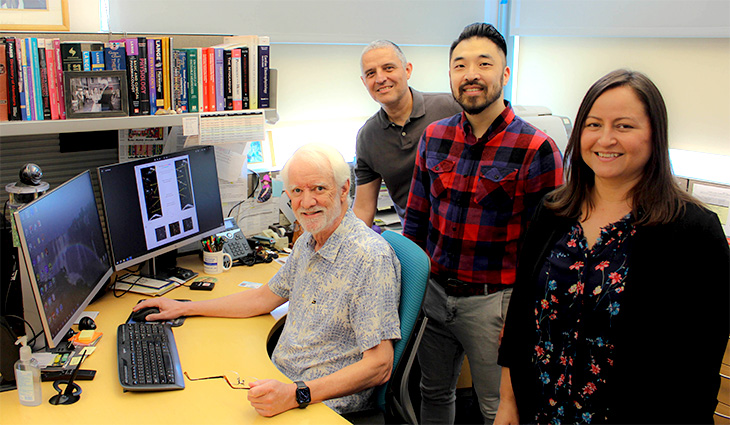
To create the new software, SparkMaster 2, Tomek worked with Bers, Ko, Manuel F. Navedo and Madeline Nieves-Cintron in the Department of Pharmacology at the UC Davis School of Medicine.
Although much of the new interface resembles the original SparkMaster, they started from scratch for the programming. Using Python, an open-source programming language that has grown in popularity, they were able to utilize many existing tools and libraries to create the unique features they wanted.

They ended up with software that can analyze calcium sparks recorded using different microscopy approaches and from cells from different tissues and genetically modified mice. Some key features of the new version include:
- improved user interface
- higher accuracy at identifying calcium release events than previous tools
- ability to identify multiple types of calcium-release events
- ability to accurately split and analyze individual sparks within spark clusters
"SparkMaster 2 is even easier to use and is much more powerful in the variety of event types it can analyze quantitatively — sparks, waves, mini-waves and latencies. And it has higher accuracy and sensitivity, resulting in fewer missed events and fewer erroneous positives," Bers said.
Bers anticipates the primary users of the software will be scientists who study calcium in muscle. “But it may be useful for researchers who study other cell types, such as neurons, that exhibit local calcium events that are important in regulating cellular function," Bers said.
Ko, who is a co-developer of the new tool, is also interested in the broader research applications for the new software.
"I'm particularly excited about the new scientific questions that SparkMaster 2 will enable the biomedical research community to answer and, in turn, the ability to more completely and accurately understand the biology that underlies human physiology in health and disease," Ko said.
Additional authors on the paper include Nieves-Cintron and Navedo from UC Davis. They were instrumental in demonstrating how the software can be used outside cardiac research, and across distinct cell types, showing its wide range of possible uses.
Funding: The project was funded by the NIH grants R01-HL142282, R01-HL092097,
P01-HL141084 (Bers), R01-HL121059, and R01-HL161872 (Navedo). J. Tomek is supported by the Sir Henry Wellcome Fellowship (222781/Z/21/Z). This research was funded in whole, or in part, by the Wellcome Trust (222781/Z/21/Z).
Resources